Notes on how to use plink¶
I found PLINK is really fast.
module load plink/2.0
subset by individuals¶
My sample ID is named like NA06985_NA06985, so the keep list just need one column.
[yli11@noderome203 test]$ head 2.list
NA06985_NA06985
[yli11@noderome203 test]$ plink2 --vcf chrAll.95samples.withinRegions.v2.03282023.vcf --keep 2.list --out test --export vcf
[yli11@noderome203 test]$ head 3.list
NA06985
NA06986
[yli11@noderome203 test]$ plink2 --vcf $f --keep 3.list --out test2 --export vcf
create allele frequency report¶
The goal is to filter out all variants that are 0/0 in your set of individuals.
plink2 --vcf $f --out test3 --freq --set-missing-var-ids @_#_\$r_\$a --new-id-max-allele-len 94
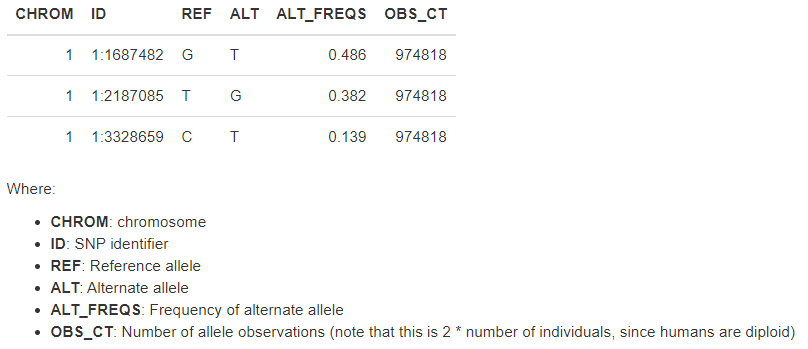
The output is test3.afreq
https://2cjenn.github.io/PRS_Pipeline/
subset by region¶
plink2 --vcf $f --extract range set1.bed --out test4 --export vcf
subset by variant ID¶
plink2 --vcf $f --extract significant.ID.list --out $f.sig --export vcf
I once had an issue getting subset, because the binary plink file uses double id. What we need to do first is to check if this is the case by generating the pgen file. If so, the id.list need to have space seprated id for two columns.
plink2 --bfile 1000G.allPops.GRCh38.allSamples.chrALL.recodeXrsid --make-pgen --out test
for the genetic variantion project, to get vcf files for the 95 donors¶
module load python/2.7.13
run_lsf.py -f /home/yli11/HemTools/share/misc/1kg.filelist --sample_list donor.list -p subset_1kg_by_people
## once it is finished, the final.vcf is what you need
Extract meta info¶
bcftools query -f '%CHROM %POS %AN_EAS\n' chrAll.95samples.withinRegions.v2.03282023.vcf.gz
[yli11@noderome153 SNP152]$ bcftools query -f '%CHROM %POS %ID %REF %ALT %GENEINFO %FREQ %COMMON\n' GCF_000001405.38.bgz
#view header
bcftools view -h GCF_000001405.38.bgz
Update SNP ID with RS ID¶
https://www.biostars.org/p/171557/
bcftools query -f ‘%ID %AF_EAS %AF_AMR %AF_EUR %AF_AFR %AF_SAS %AF_EUR_unrel %AF_EAS_unrel %AF_AMR_unrel %AF_SAS_unrel %AF_AFR_unrel %MAF_EUR_unrel %MAF_EAS_unrel %MAF_AMR_unrel %MAF_SAS_unrel %MAF_AFR_unreln’ ${COL1} > $label.AF.txt
intersect bed file¶
PLINK only work for vcf file with genotypes, for gnomad vcf file, use vcftools for region extraction:
vcftools --gzvcf gnomad.genomes.v4.0.sites.chrY.vcf.bgz --bed 1115_OT_flank100.bed --out output_prefix --recode --keep-INFO-all